Alanine Dodecapeptide Folding
SPONGE 1.4 tutorial.This page was translated by GPT-5.5 AI.
Alanine Dodecapeptide Folding
Last updated:
2024/01/01
Main topics:
- Build an implicit-solvent model of alanine dodecapeptide with Xponge
- Run the simulation with SPONGE 1.4
- Visualize the results with VMD
Software Installation
See Installation on Linux and Installation on Windows.
Model Building
Open a terminal and start Python:
python
In the Python terminal, enter the following Python commands in order.
1. Import the Required Modules
import Xponge #Xponge itself
import Xponge.forcefield.amber.ff19sb #The force field used
from Xponge.forcefield.special import gb #The implicit-solvent GB model used
2. Obtain the Xponge Molecule Instance
mol = NALA + ALA * 10 + CALA
3. Set the GB Parameters
gb.Set_GB_Radius(mol)
4. Save the Input Files
Save_PDB(mol, "ALA.pdb")
Save_SPONGE_Input(mol, "ALA")
Molecular Dynamics Simulation
In the working folder, create an mdin.txt file with your preferred text editor:
ALA12 simulation
pbc = 0
mode = NVT
dt = 2e-3
cutoff = 999
constrain_mode = SHAKE
step_limit = 5000000
thermostat = middle_langevin
default_in_file_prefix = ALA
For the meanings of these commands, see Input Commands.
Then enter the following command in the terminal:
SPONGE -mdin mdin.txt
Analysis and Discussion
Result Analysis
You can read the generated mdout.txt file to obtain the energy data. Use the following Python script to read the energy changes:
from Xponge.analysis import MdoutReader
mdout = MdoutReader("mdout.txt")
import matplotlib.pyplot as plt
plt.plot(mdout.time, mdout.potential)
plt.show()
The resulting plot is shown below:
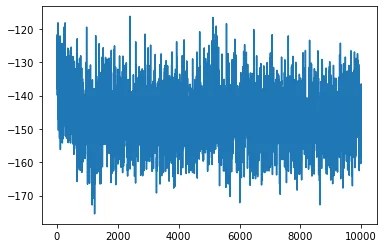
You can also use VMD to visualize the trajectory:
vmd -sponge_mass ./ALA_mass.txt mdcrd.dat

