Polymer Modeling
SPONGE 1.4 tutorial.This page was translated by GPT-5.5 AI.
Polymer Modeling
Updated
2023/04/27
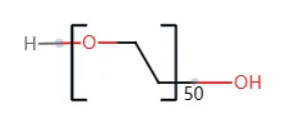
This tutorial uses the construction of a polyethylene glycol molecule with 50 repeating units, shown below, as an example.
Building a polyethylene glycol molecule requires the following four residue types:
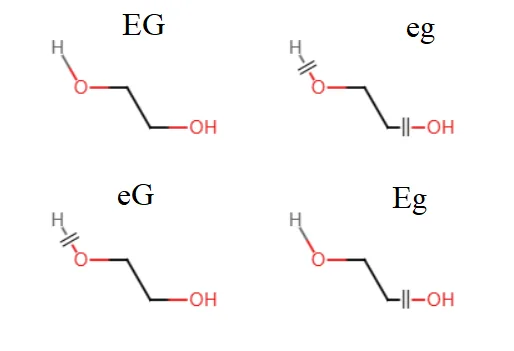
Here, EG, eg, eG, and Eg are arbitrary names that do not conflict with names in your existing force field. EG is the name used in this tutorial; it is an abbreviation for Ethylene glycol. E indicates that the head hydrogen is present, while e indicates that the head hydrogen is absent. G indicates that the tail hydroxyl group is present, while g indicates that the tail hydroxyl group is absent.
The final goal is to build the following molecule:
mol = Eg + eg * 48 + eG
That is, the molecule shown below:
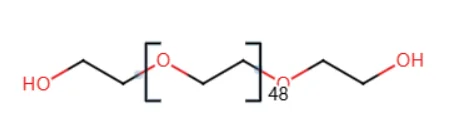
1. Build the Force Field for the EG Monomer
import Xponge
import Xponge.forcefield.amber.gaff as gaff
"""Get an assignment from the SMILES of ethylene glycol so that force-field terms can be assigned."""
assign = Xponge.Get_Assignment_From_Smiles("OCCO")
"""Get the corresponding atom types and charges."""
assign.determine_atom_type("gaff")
"""If high accuracy is required, use resp charges.
assign.calculate_charge("resp")
If high accuracy is not required, tpacm4 charges can also be used."""
assign.calculate_charge("tpacm4")
"""Convert the assignment to a ResidueType."""
EG = assign.to_residuetype("EG")
"""Save the file."""
save_mol2(EG, "EG.mol2")
Open the EG.mol2 file to inspect the atom types, charges, and related information.
@<TRIPOS>MOLECULE
EG
10 9 1 0 1
SMALL
USER_CHARGES
@<TRIPOS>ATOM
1 O -1.711 0.268 -0.387 oh 1 EG -0.633105
2 C -0.485 -0.387 -0.314 c3 1 EG 0.105505
3 C1 0.603 0.581 0.097 c3 1 EG 0.105505
4 O1 1.843 -0.049 0.177 oh 1 EG -0.633105
5 H -2.186 0.250 0.500 ho 1 EG 0.405197
6 H1 -0.289 -0.876 -1.280 h1 1 EG 0.061202
7 H2 -0.563 -1.156 0.505 h1 1 EG 0.061202
8 H3 0.320 0.955 1.112 h1 1 EG 0.061202
9 H4 0.701 1.428 -0.585 h1 1 EG 0.061202
10 H5 1.767 -1.015 0.175 ho 1 EG 0.405197
@<TRIPOS>BOND
1 1 2 1
2 1 5 1
3 2 3 1
4 2 6 1
5 2 7 1
6 3 4 1
7 3 8 1
8 3 9 1
9 4 10 1
@<TRIPOS>SUBSTRUCTURE
1 EG 1 **** 0 **** ****
From the file, we can see that the hydrogen atom named H is bonded to the first hydroxyl oxygen. The later hydroxyl atoms are named O1 and H5. These two pieces of information will be used to generate the other residues.
2. Build the Force Fields for Residues eg, Eg, and eG
"""Continue from the monomer construction above."""
"""Build residue eG."""
eG = EG.deepcopy("eG")
"""The first argument of Omit_Atoms is the atom to delete, and the second argument is the charge of the residue after deletion.
Here, one hydrogen ion is deleted, so the residue charge should be -1."""
eG.Omit_Atoms([eG.H], charge=-1)
save_mol2(eG, "EG1.mol2")
"""Repeat the process above to build residue Eg."""
Eg = EG.deepcopy("Eg")
Eg.Omit_Atoms([Eg.O1, Eg.H5], charge=+1)
save_mol2(Eg, "EG2.mol2")
"""Repeat the process above to build residue eg."""
eg = EG.deepcopy("eg")
eg.Omit_Atoms([eg.H, eg.O1, eg.H5], charge=0)
save_mol2(eg, "EG3.mol2")
3. Connect the Residues
"""Continue from the construction process above."""
"""The head atom name of residue eg is O.
The atom adjacent to the head atom on the main chain of residue eg is C.
The bond length at the head atom of residue eg is 1.2 angstroms."""
eg.head = "O"
eg.head_next = "C"
eg.head_length = 1.2
"""The tail atom name of residue eg is C1.
The atom adjacent to the tail atom on the main chain of residue eg is C.
The bond length at the tail atom of residue eg is 1.2 angstroms."""
eg.tail = "C1"
eg.tail_next = "C"
eg.tail_length = 1.2
"""Set the connection rules for bond angles and dihedral angles below.
O and C of eg form a 109.5-degree bond angle with the tail atom of another residue.
C and C1 of eg form a 109.5-degree bond angle with the head atom of another residue.
C1, C, and O of eg form a -180-degree dihedral angle with the tail atom of another residue.
O, C, and C1 of eg form a -180-degree dihedral angle with the head atom of another residue."""
PI = 3.141592654
eg.head_link_conditions.append({"atoms": ["C", "O"], "parameter": 109.5 / 180 * PI})
eg.tail_link_conditions.append({"atoms": ["C", "C1"], "parameter": 109.5 / 180 * PI})
eg.head_link_conditions.append({"atoms": ["C1", "C", "O"], "parameter": -PI})
eg.tail_link_conditions.append({"atoms": ["O", "C", "C1"], "parameter": -PI})
"""Get the molecule."""
mol = eg * 50
"""Convert the first residue to Eg.
Convert the last residue to eG.
Fill in the atoms of Eg and eG."""
mol.residues[0].set_type("Eg")
mol.residues[-1].set_type("eG")
mol.add_missing_atoms()
"""Use gaff parmchk2 to find missing force-field parameters."""
gaff.parmchk2_gaff(mol, "eg.frcmod")
"""Save the files."""
save_mol2(mol, "PEG.mol2")
save_sponge_input(mol, "PEG")
4. Inspect the Model and Run a Molecular Dynamics Simulation
Use vmd to inspect the final generated PEG.mol2. It should look like this:

5. Notes
-
This example covers only the modeling of the polyethylene glycol part. Additional processing, such as solvation, is the same as in ordinary workflows.
-
The dihedral angles and other parameters in this example can be adjusted according to your own needs.
-
This example assigns force-field parameters using only the ethylene glycol monomer. You can also assign force-field parameters using triethylene glycol; the difference lies in which atoms are omitted by
omit_atoms. -
If you want to build an infinitely long molecule, do not call
set_typeat the end. Instead, connect the head and tail of the zeroth and last residues inmol, as shown below:"""Get the molecule.""" mol = eg * 50 """Convert the first residue to Eg. Convert the last residue to eG. Fill in the atoms of Eg and eG. mol.residues[0].set_type("Eg") mol.residues[-1].set_type("eG") mol.add_missing_atoms()""" mol.add_residue_link(mol.residues[0].C1, mol.residues[-1].O)
